Years of laboratory experiments with microorganisms have revealed how cells are capable of adapting to new challenges over short evolutionary timescales. However, what dictates which mutations will contribute to a complex adaptation, and what aspect of cell biology they alter, has remained elusive. To investigate these points, Fumasoni and Murray took advantage of a previously characterized adaptation to constitutive DNA replication stress to study how the genomic features of cells affect their evolution.
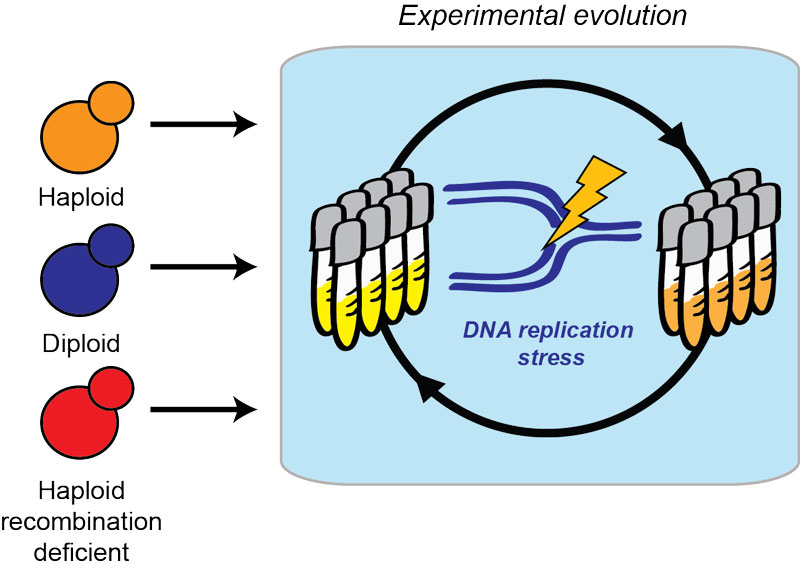
Figure: Experimental design with S. cerevisiae cells
DNA replication stress is described as a wide variety of insults, either genetic or environmental, that perturb DNA synthesis. Genetic changes that produce replication stress are frequent in cancer. In an earlier study the authors evolved haploid budding yeast cells experiencing constitutive DNA replication stress and showed how the adaptation to this stress was achieved through mutations altering three distinct cellular processes: sister chromatid cohesion, DNA replication and cell cycle progression.
In a new study, the authors exploited the high reproducibility of this laboratory evolution experiment to test the possibility that particular features of the cells’ genomes could alter how they adapt to DNA replication stress. To do this, Fumasoni and Murray applied the same selective pressure (DNA replication stress) to diploids (which carry an extra copy of the genome compared to haploids) and recombination deficient cells (which have difficulties in using recombination between identical DNA sequences to repair DNA damage). Despite concerns that these genomic features could have delayed or impaired cell adaptation, the authors found that adaptation in diploids and cells that lack DNA recombination happened as fast, or faster than in the original haploid cells. Through a combination of genome sequencing and engineering, Fumasoni and Murray also showed that the genes mutated, but not the cellular process they contributed to, were different in the strains with different genomic features.
In the new study, published in PLOS Genetics, the authors propose that a universal evolutionary solution to DNA replication stress exists, but that its precise implementation through mutations is shaped by the genomic features of the cells subjected to the stress. This knowledge increases our understanding of how cells adapt to perturbations in their core machineries, and thus improves our ability to predict similar adaptive processes outside the laboratory, including those that occur in the genesis and evolution of cancer.
Marco Fumasoni, former postdoc in the Murray lab at Harvard University, is now a group leader at the Instituto Gulbenkian de Ciência, where he continues to investigate the interplay between genome maintenance mechanisms and evolutionary forces in shaping cellular features.
Press release from Gulbenkian Science
|
HFSP award information Long-Term Fellowship (LT000786/2016-L): Experimentally evolving biological novelty during adaptive radiation of budding yeast Fellow: Marco Fumasoni |


































