Our HFSP project aims to get structural scattering information from individual molecules rather than from crystals. The basis of the structural information is X-ray scattering, particularly from a XFEL (X-ray Free Electron Laser) source. This is a coherent source of X-rays, but useful structural information requires sufficient scattering from each sample. At this stage of research, a good candidate for such a sample is DNA origami. DNA origami is a large DNA complex containing about 7200 base pairs, formed into a particular shape from a viral single stranded genome and 200-250 smaller 'staple strands'. For our purposes, we are actually trying to combine more than a single DNA origami molecule to act as a sample, origami molecules that can have the same or different shapes.
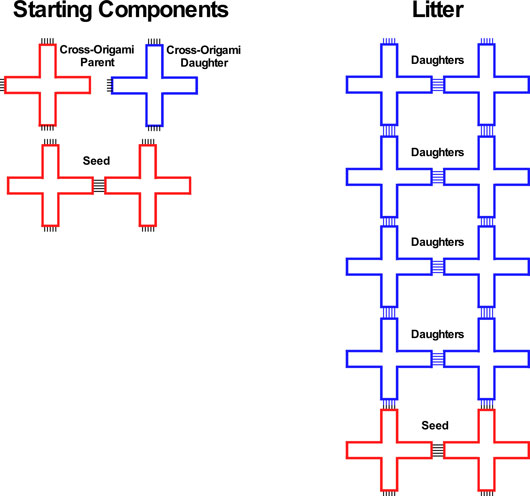
Figure. Formation of litters: On the left the components of the DNA cross-origami seed tile are shown in red, and a complete seed is shown as a double red tile. The components of the offspring are shown in blue. Note that the tile can be rotated in the plane. The small black lines represent half-complementary or paired strands that can (top) or do cohere (in the seed). The right side shows the production of a blue litter from a group of blue tiles, catalyzed by a red seed. Note that the seed can be in the middle of the array, as well as at one end.
The individual DNA origami molecules can be used in self-replication schemes, as well as providing samples for scattering studies. This has been done previously, where a parent or seed species pairs with potential daughters, to produce a second copy of itself. This is a one-sided interaction, so that each parent leads to a single daughter in the next generation. This interaction, related, but not identical to that used in the scattering experiments, consists of one face of one origami talking to two daughters, which are then fused together by cross-linking and then released by heating to provide parents for the next generation.
In the work described in the Proc. Nat. Acad. Sci. article (Zhuo et al.), we found that the use of parent/seed origami tiles with multiple binding surfaces can produce 'litters' of progeny for the next generation. These particular origami tiles are cross-shaped, so that opposite faces of the cross can be made to recognize daughter components. The work described shows a small 'crystalline' intermediate, which is then released to produce the litter of daughters. If we do something similar with the multiple-origami-tile samples we are using for our diffraction studies, we can alter the protocol to yield cross-linked multiple copies of the sample of interest. This procedure will enable us to increase the scattering from the sample of interest if it is not scattering adequately for a good signal.


































