To better understand how SARS-CoV-2 infects the lungs and causes disease, the team of Professor Young Seok Ju of the Graduate School of Medical Science and Engineering at KAIST, in collaboration with the Wellcome-MRC Cambridge Stem Cell Institute, turned to organoids – ‘mini-organs’ grown in three dimensions to mimic the behaviour of tissue and organs.
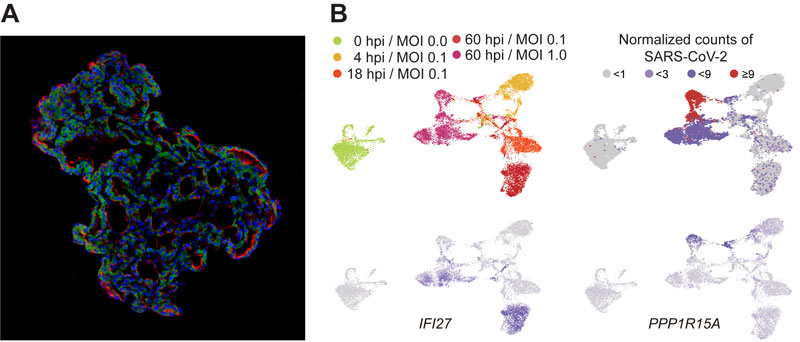
Figure: (A) Representative image of three-dimensional human lung alveolar models showing alveolar stem cell marker, HTII-280 (red), and SARS-CoV-2 entry protein, ACE2 (green). (Credit: Jeonghwan Youk, Taewoo Kim, and Seon Pyo Hong).
(B) Tracing SARS-CoV-2 infection using single-cell transcriptome techniques. Top left: infected cells show different gene expressions as time goes by after infection [hpi: hours post infection]; top right: viral burden increases from the first day after infection to the full extent at 60 hpi; bottom left: genes of intrinsic immune responses (i.e. IFI27) are actively transcribed at 60 hpi; bottom right: a substantial fraction of alveolar cells at 60 hpi expresses cell death markers (i.e. PPP1R15A).
The team used human lung tissue donated from hospitals to collect stem cells of lung alveoli, or alveolar type 2 cells. By cultivating these cells in chemically defined conditions, the team was able to grow self-organising alveolar-like 3D structures that mimic the behaviour of key lung tissue. Long-term expansion of the models was possible for ~10 months in laboratory conditions.
Then, the team infected the mini-lungs with SARS-CoV-2 viral particles and studied how human lung cells responded to the virus using a combination of imaging and genomics techniques. Most interestingly, when the 3D models were exposed to SARS-CoV-2, the virus began to replicate rapidly, reaching full cellular infection just six hours after infection. Shortly after the viral replication, infected lung cells began to produce interferons – proteins that act as warning signals to neighbouring cells, telling them to activate their antiviral defences. After 48 hours, the interferons triggered an innate immune response – its first line of defence – and the cells started fighting back against infection. Sixty hours after infection, a subset of alveolar cells with insufficient cellular defence began to disintegrate, leading to loss-of function and cell death.
The model has many applications for tackling the many unanswered key questions about SARS-CoV-2, such as understanding genetic susceptibility to SARS-CoV-2, assessing relative infectivity of viral mutants, and revealing the damage processes of the virus in human alveolar cells. Most importantly, it provides the opportunity to develop and screen potential therapeutic agents against SARS-CoV-2 infection. The models are also applicable to the study of basic biology of human alveolar cells and respiratory diseases.
The research team were able to start the study immediately after the current outbreak, because human organoids (3D culture models) had been actively established and extensively investigated for conducting our HFSP project, which aims to understand mutagenesis and tumour evolution in human cells. The urgent situation of a world-wide pandemic pushed the team to apply a fraction of the cellular models in the infection studies. Although it is a serendipitous study and also an accidental development, our efforts have produced a powerful new community platform to study respiratory viruses and to explore possible treatments. Currently, the team is expanding their viral infection studies and combining them with other techniques, such as co-culture experiments with immune cells.
|
HFSP award information Research Grant - Early Career (RGY0071/2018): Tracing AID/APOBEC- and MSI-mediated hyper-mutagenesis in the clonal evolution of gastric cancer Principal investigator: Young Seok Ju, Korea Advanced Institute of Science and Technology, Daejeon, Republic of Korea |


































