Unlike physics, where many natural phenomena can be described by universal laws, biology is full of exceptions. That is precisely why biologists study many organisms. Broad comparisons reveal recurring patterns that allow a certain degree of generalization. Yet, as we widen the lens of inquiry, the “exceptions” often reappear, forcing us to rethink what we believed was a general rule. This study is one such case.
At its core, this study concerns protein evolution. Proteins are chains of amino acids, whose sequences shape their structure and function. When environments change, natural selection can favor amino-acid substitutions that alter a protein in ways that improve its function under new conditions. In some cases, two distantly related species can start out with completely different amino acids at important functional sites in the protein, and yet independently acquire the same substitution at the same site over evolutionary time. This kind of repeated, site-specific change is molecular convergence. These changes cannot be attributed to random chance. Instead, it reflects adaptation acting on the available set of amino acids to reach the same molecular “solution” despite different evolutionary histories.
Over the decades, researchers have found that certain classes of genes, such as those encoding for proteins involved in gas transport and sensory perception, are frequent targets of adaptive change. In contrast, pleiotropic genes have traditionally been viewed as poor candidates for adaptation. Pleiotropy refers to a single gene influencing multiple biological systems. Because changes to such genes can have widespread effects, a substitution that is beneficial in one context may be harmful in another, leading pleiotropic proteins to be strongly constrained and conserved. But, as with many ideas in biology, the more systems we examine, the more we encounter exceptions.
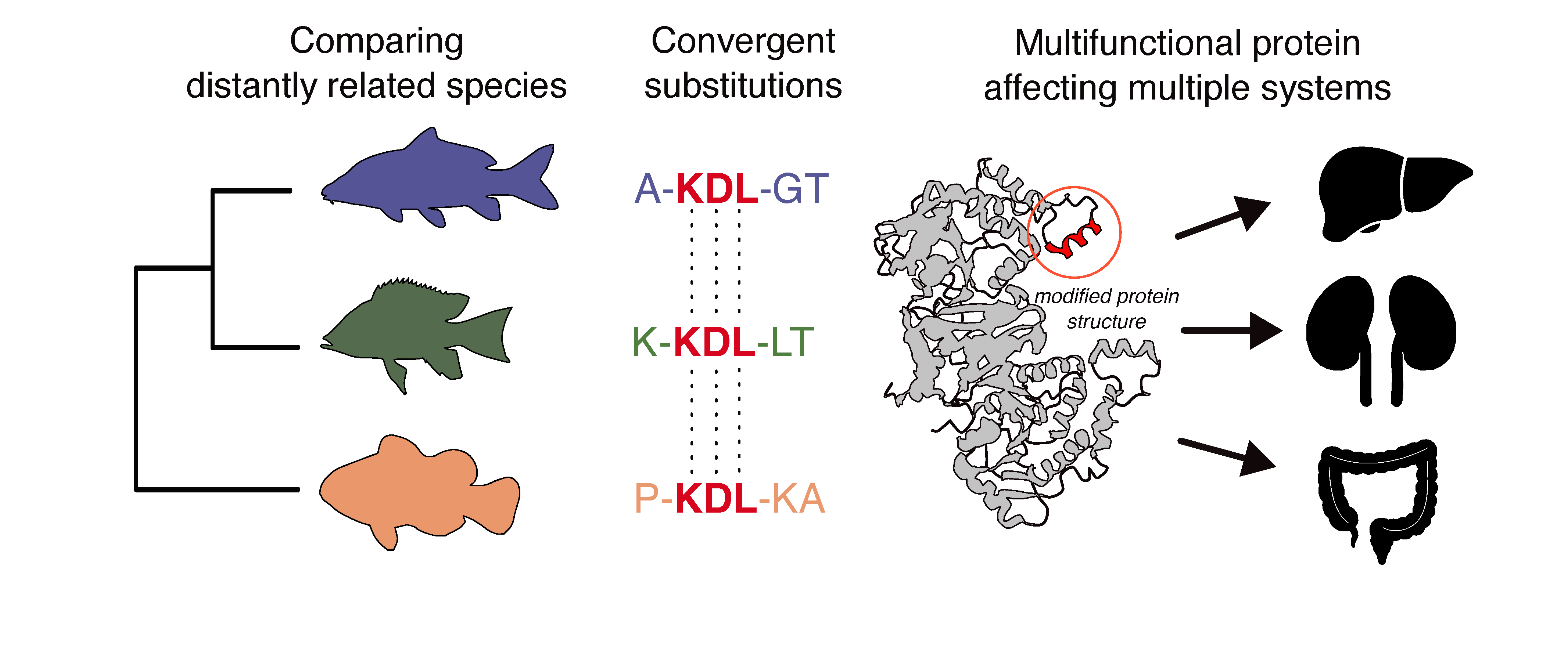
In this study, HFSP Long-Term Fellowship awardee Agneesh Barua analyzed over two million protein-coding genes from 143 teleost fish genomes to search for signatures of molecular convergence. He used a novel, unsupervised, data-driven approach to identify convergent signals without relying on a pre-defined set of genes or pathways. This approach yielded 89 gene families with evidence of molecular convergence. Surprisingly, a majority of these genes showed broad expression across tissues and cell types. Moreover, genetic perturbations of these genes were associated with phenotypic effects spanning multiple biological systems. Together, these lines of evidence point to their pleiotropic nature and suggest that adaptive variation can, under some circumstances, accumulate even in pleiotropic genes.
Using targeted simulations, Barua and his host supervisor, Marc Robinson-Rechavi at the University of Lausanne, further showed that in specific evolutionary scenarios, adaptation via pleiotropic genes may not only be possible, but even favored. While this study is not the first to raise this possibility, it is the first to demonstrate the pattern at such a broad phylogenetic scale in an exceptionally diverse vertebrate lineage like teleost fishes. As comparative genomics has expanded, similar observations have become increasingly difficult to ignore. Researchers are now at a point where the classic view that pleiotropic genes are generally off-limits to adaptation needs refinement in light of accumulating evidence.


































