Agricultural and wild plants host diverse microbes on and in their leaves. Amongst these microbial consortia are numerous pathogens. While pathogens are common colonizers of plants, most plants manage to stay healthy. What prevents common pathogens from killing their plant hosts? Plant genetics definitely plays a role—decades of plant pathology work have revealed plant genes that confer general resistance and also specialized resistance to specific pathogens. However, plants of the same genotype can also show different susceptibility to pathogens, leading to the question: how do surrounding commensal microbes influence the spread of a pathogen?
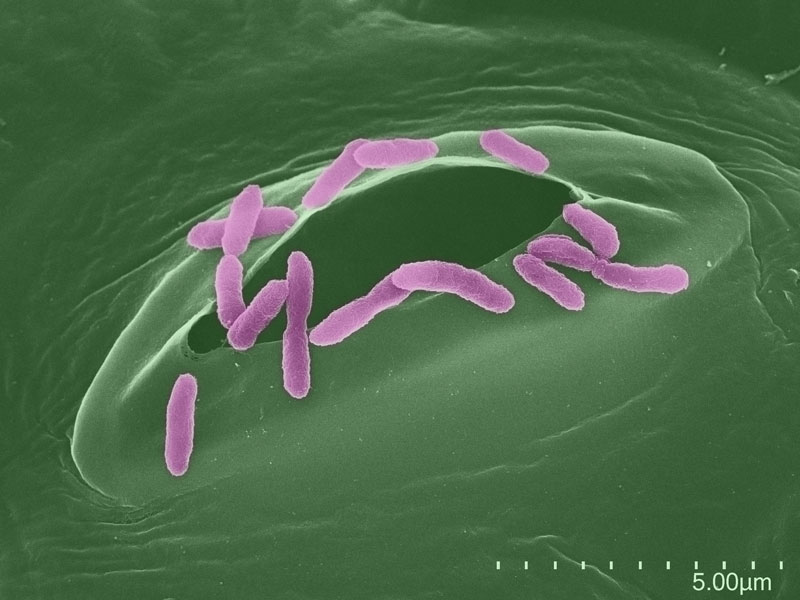
Image: Pseudomonas cells surrounding a stoma of an A. thaliana plant. Credit: Jürgen Berger, Detlef Weigel, Sonja Kersten
In a project led by Dr. Or Shalev involving HFSP fellows Dr. Talia Karasov and Dr. Derek Lundberg, a team of researchers in the lab of Dr. Detlef Weigel at the Max Planck Institute (MPI) for Biology in Tübingen, Germany, sought to understand how the presence of surrounding commensal bacteria influenced disease progression by bacterial pathogens. Working on the model plant Arabidopsis thaliana, the team treated plants with different combinations of pathogenic and commensal Pseudomonas bacteria. The Pseudomonas strains were genetically barcoded with a unique DNA sequence before mixing the combinations, enabling all strains to be easily and unambiguously distinguished. The team then tracked the abundance of the diverse pathogenic and commensal bacterial strains while simultaneously monitoring plant health. They found that the commensal microbes often suppressed the proliferation and disease development caused by pathogenic bacteria, although not all plant genotypes benefited from the suppressive ability of the commensal microbes.
Agricultural pathogens are a significant cause of global food insecurity, and finding ways to combat pathogen spread remains a central challenge, particularly as climate change expands the range of many pathogens to higher latitudes. This study supports a role for surrounding microbes in combating disease. Ultimately, harnessing the plant microbiome to combat disease has potential to increase agricultural yields with minimal environmental impact.
|
HFSP award information Long-Term Fellowship (LT000348/2016-L): The population genomics of pathogen spread in natural Arabidopsis thaliana populations Fellow: Talia Karasov |
|
HFSP award information Long-Term Fellowship (LT000565/2015-L): Evolutionary genetics of plant-microbe interactions in natural populations Fellow: Derek Lundberg |


































